PepKit Documentation
PepKit
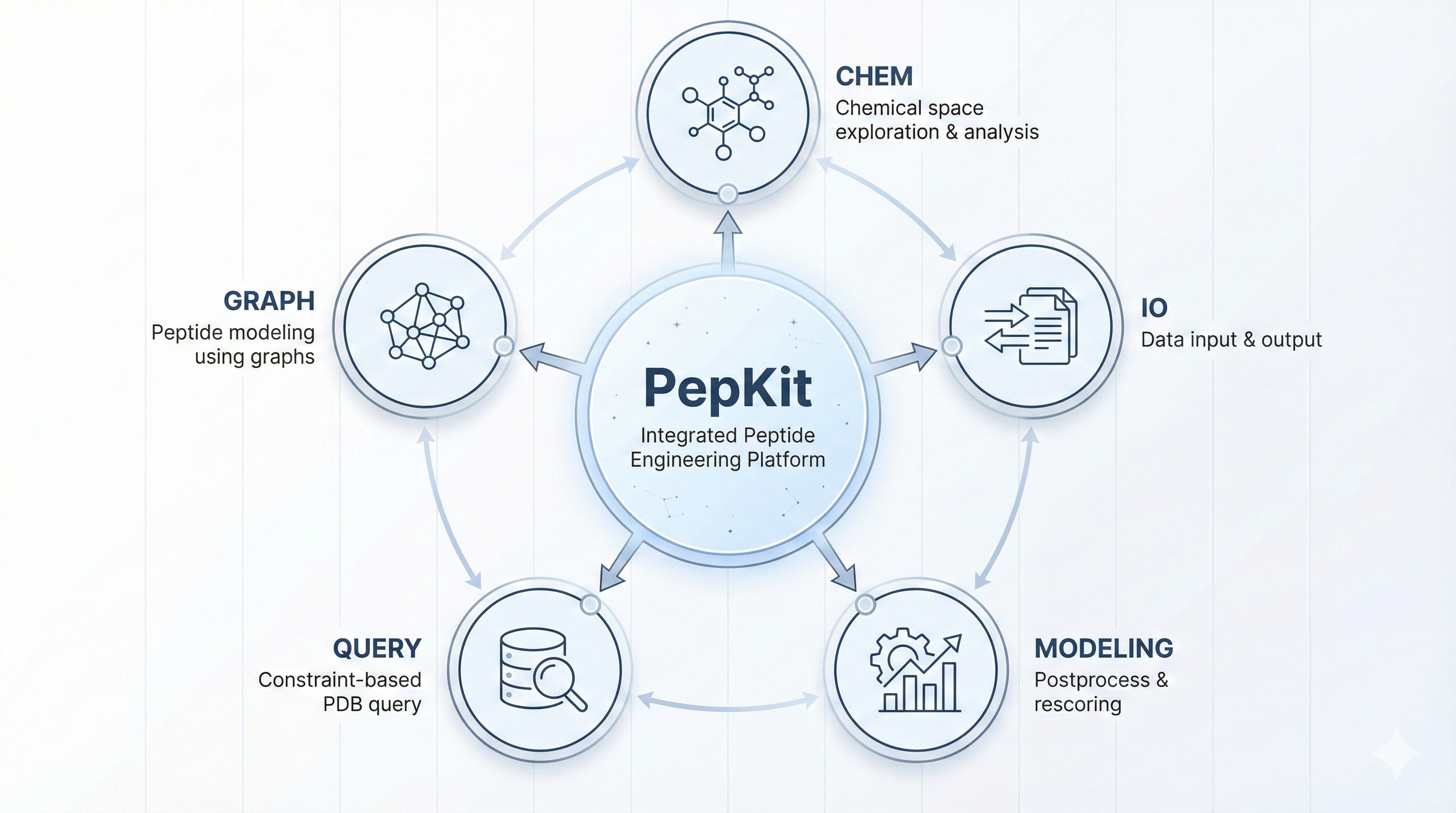
A modular toolkit for peptide chemistry, graph-based modeling, PDB queries, and structure rescoring.
At a glance
What it helps with
Peptide format conversion, dataset standardization, feature extraction, graph-based modeling, PDB queries, and structure-level post-processing and rescoring.
Who it is for
Users building peptide datasets, exploring graph-based peptide representations, querying peptide–protein complexes, or benchmarking structural models.
Quick install
pip install pepkit
Minimal example and full quickstart are in the Getting Started guide.
Start here
Getting Started
Install PepKit and run a minimal end-to-end example.
Open guide →Chem
FASTA ⇄ SMILES conversion, standardization, and descriptors.
Explore chem →Graph
Peptide modeling using graph-based representations and utilities.
Explore graph →Query
Constraint-based PDB queries for peptide–protein complexes.
Explore queries →Modeling
Post-process and rescore predicted complexes (e.g. AlphaFold outputs).
Explore modeling →API Reference
Programmatic reference for PepKit modules and functions.
Explore API →